
Genomics platform services
Services
Molecular biology
Molecular Biology Techniques provide opportunity to study the composition, structure and interactions of cellular molecules. These techniques include DNA cloning, bacterial transformation, transfection, chromosome integration, cellular screening, Nucleic acid extraction, Nucleic acid amplifications and synthesis (PCR and cDNA synthesis) DNA sequencing, molecular hybridisation, random mutagenesis, among others. We offer a portfolio of the most essential techniques as well as many custom molecular engineering technologies.
Library preparations
Library preparation forms the basic initial stage of NGS. Prior to sequencing nucleic acid from samples, DNA or RNA must be extracted, fragmented, end-repaired, and covalently attached to adapters using ligation or tagmentation processes.
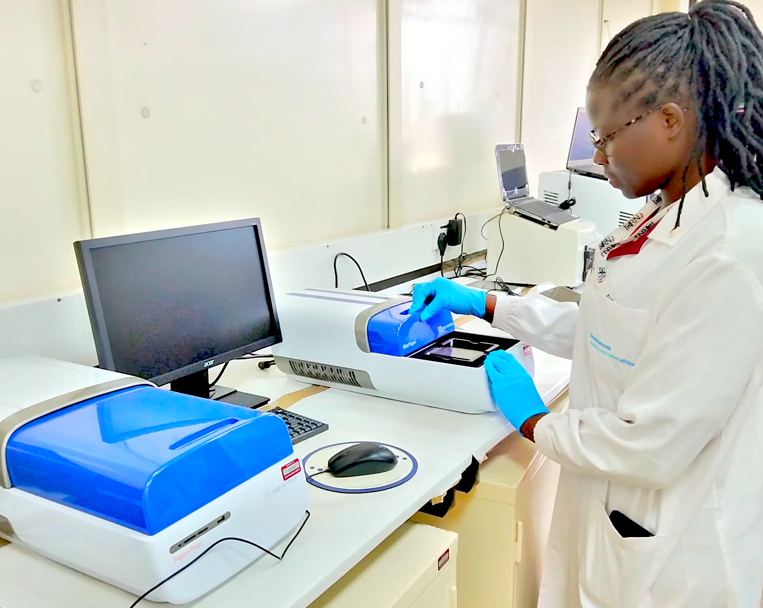

The standard and quality of NGS library preparation determines the accuracy of NGS data. We offer a arrange of NGS library preparation tools for high quality DNA or RNA libraries for whole genome sequencing (WGS), whole transcriptome analysis (WTA), total RNA sequencing, and miRNA and small RNA sequencing applications. We prepare sequencing libraries for both Illumina and Oxford Nanopore technologies. Several equipment are used in library preparation stage of next-generation sequencing.

RT-qPCR analysis
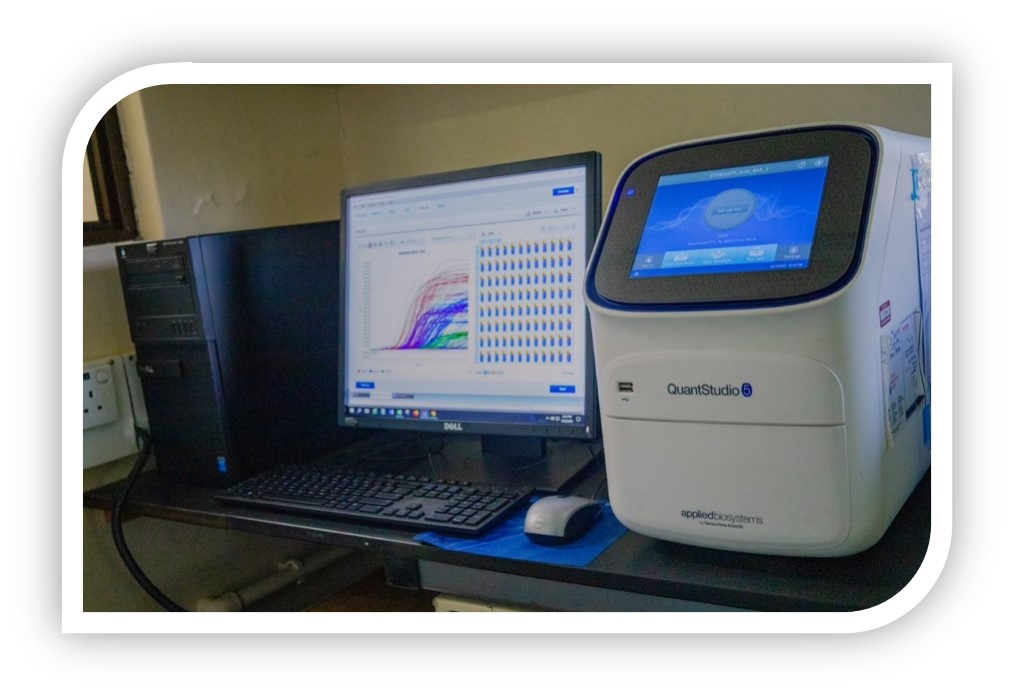
qPCR and RT-qPCR are now widely used in a range of applications such as gene expression analysis, SNP identification, genotyping, contamination screening, and molecular diagnostics for infectious diseases. An optimised protocol with standardised step by step workflow is critical for reliable and reproducible results for these quantitative amplification techniques. RT-qPCR involves RNA quantitation, in which DNA amplification by PCR is preceded by a reverse transcription (RT) step to generate cDNA from RNA in either a two-step RT-qPCR process (cDNA synthesis separated from amplification) or as an upfront step within the same reaction (one-step RT-qPCR). Our platform has both technical and infrastructural capacity to conduct qPCR experiments in high-throughput formats.
Our gene expression studies are versatile and expandable to include more targets, multiple controls, and additional replicates; screen at larger scale; and diagnostic tests can accommodate many samples. Our instruments have automated liquid handling systems thus eliminate error, increase efficiency, and minimise manual setup as a workflow bottleneck.

Nucleic acid extractions
Nucleic acid extraction processes include isolation, purification, concentration and quantification. Within our platform we have developed protocols for RNA and DNA extraction for a wide range of nucleic acid extraction kits. We perform extraction using high throughput automatic extraction machines such as Tanbead (Fig. 9), semi-automatic processes with column-based technologies and also manual (gold standard) lysis and precipitation techniques described in Maniatis laboratory manuals.
Single-Cell Gene Expression
In multicellular organism’s various cell types within the same population play distinct functions and comprise subunits with unique transcriptional profiles. However, genomic studies were previously focused on measurements for whole tissues and as such limited to average expression profile for all the constituent cells. In multicellular organisms’ different cell types within the same population can have distinct roles and form subpopulations with different transcriptional profiles.
Through the Single gene expression services, we offer in-depth gene expression to the subpopulation
levels, link changes in the expression profile to specific regulation or compositional, and categorise cellular progression through differentiation expression profiles in time and at various cell development stages. The genomic capabilities under this platform offer high resolution assessment of transcriptomes of individual cells to elucidate cell-to-cell differences. Evaluate the distinct biology of individual cells in complex tissues and reveal responses of cellular subpopulations to candidate vaccines (Immunogenetics) or other environmental changes under investigation.
Whole-Exome Sequencing
This service identifies coding variants across wide range of applications including population genetics.
Shotgun Metagenomics
A genomic technology that allows for a detailed assessment of all organisms within a complex sample. The technology uses genetic approaches to detect, determine and characterise microorganisms present in a complex sample like infected tissues, swabs or environmental samples (like wastewater, soil of faecal material). The technology can be used as diagnostic tool for infectious diseases by using the detected microorganisms to make etiological inferences. This technology is critical for characterizing unculturable microorganisms which are ordinarily difficult to analyse following classical microbiology approaches.
Microbial Whole-Genome resequencing
Targets sequencing services for the entire genome of a bacteria, viruses and other microbes (≤ 5 Mb) and alignment to a known reference genomes to determine variation (Isolate/variant/strain detection). Small microbial genomes sequencing support food safety testing for public health, infectious disease surveillance, molecular epidemiology studies, and environmental metagenomics.
Through the microbial Sequencing we:
- Identify circulating variants through genomic surveillance
- characterise genes from single organism isolates
- Perform detailed genomic analysis of the microbial or viral populations
- Support discovery of new biomarkers in a microbial/viral samples




