Scientists identify genomic regions controlling important genetic traits for Napier grass
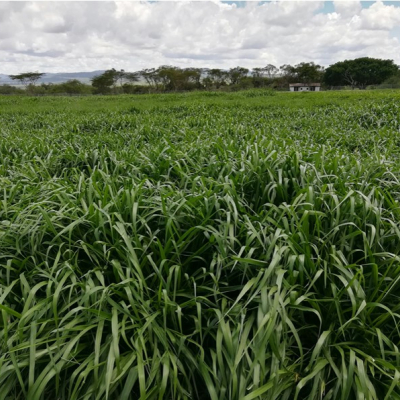
Napier grass, also known as elephant grass, is one of the most important fodder crops grown across the global tropics. As this new year commences, a team of scientists from East Africa, China and South Korea have identified 74 genomic regions containing quantitative trait loci (QTL) – sections of DNA which correlate with a broad range of morphological, agronomic, and feed quality traits.
In the study which is published in the journal Frontiers in Plant Science, the researchers applied several different approaches to reduce environmental variation, including spatial analysis of phenotype - observable trait - data, in order to accurately determine the genetic responses of Napier grass genotypes to various treatments.
Their work was enhanced by the fact that many of the molecular markers identified could be mapped onto the recently assembled Napier grass genome. This greatly improved the researchers’ abilities to identify the genomic regions and QTL that control important forage traits. These can be used in the future for genetic improvement of Napier grass varieties.
Meki Shehabu Muktar, lead author of the paper and a scientist at the International Livestock Research Institute (ILRI) in Ethiopia, said, ‘This study has identified two major QTL associated with anthocyanin pigmentation. Anthocyanins have a role in abiotic and biotic stress tolerance and may impact animal nutrition and greenhouse gas mitigation.’
He added that future research would focus on these topics.
Jorge Fernando Pereira, a researcher at the Brazilian Agricultural Research Corporation (EMBRAPA) which works closely with forage researchers at ILRI, shared, ‘We are very excited by the results. These results open up a vast range of possibilities for studying and applying the QTL and markers found to be associated with important plant features such as drought tolerance and agro-morphological traits.’
He further added that he hopes scientists and breeders will explore more of these QTL and markers to speed up the breeding cycle which would increase genetic gains, especially for complex traits.
Chris Jones, who led the study at ILRI, said, ‘I am most proud that we have identified QTL associated with a whole range of morphological, agronomic, and feed quality traits. I am particularly excited to see how the plant breeding capabilities at EMBRAPA and elsewhere can make use of these resources to support the development of new and improved varieties which will be used to increase livestock production.’
This research was supported by the CGIAR Research Program (CRP) on Livestock, the Rural Development Administration (RDA) of South Korea, the Federal Ministry for Economic Cooperation and Development (BMZ), and Guangxi University of Science and Technology in China.
Read the full publication: Insights into the Genetic Architecture of Complex Traits in Napier Grass (Cenchrus purpureus) and QTL Regions Governing Forage Biomass Yield, Water Use Efficiency and Feed Quality Traits
For more information on Napier grass varieties contact Chris Jones (ILRI) and Jiyu Zhang (Lanzhou University).
Related publications and additional information
- Yan Q, Wu F, Xu P, Li J, Chen D, Sun Z, Gao L, Lu L, Muktar M, Jones C, Yi X and Zhang J. (2020). The elephant grass (Cenchrus purpureus) genome provides insights into anthocyanidin accumulation and fast growth. Molecular Ecology Resources
- Simeão RM, Resende MDV, Alves RS, Pessoa-Filho M, Azevedo ALS, Jones CS, Pereira JF and Machado JC. (2021). Genomic Selection in Tropical Forage Grasses: Current Status and Future Applications, Frontiers in Plant Science, 12: 761.
- A new study gives insights on Napier (elephant) grass, a fast-growing tropical grass.
Photo credit: Man farming Napier grass (ILRI/Melkamu)