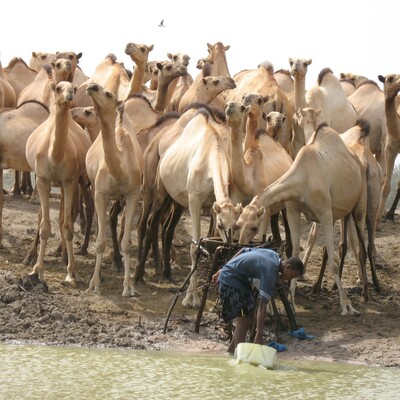

Samuel Oyola is ILRI’s Specialist Scientist in Genomics and Molecular Biology. He holds a PhD in Molecular and Cellular Biology from the University of Cambridge. Before joining ILRI, he studied functional genomics of Leishmaniasis and host-parasite interaction as a postdoctoral fellow at the University of York. He then took a Scientist position at the Wellcome Trust Sanger Institute, Cambridge UK, where he worked on malaria; developing and applying high throughput genomic technologies to study natural genetic variations in malaria parasite populations. He developed novel molecular tools that enable application and translation of genomic technologies into basic healthcare and public health applications. At ILRI, Samuel is using his experience and expertise in modern genomics, biotechnology and molecular biology to study several aspects of animal and human health. Under epidemiology, Samuel is using modern genomic and bioinformatic tools to study epidemiology of rift valley fever virus (RVF), African swine fever virus (ASFV), Newcastle disease virus (NDV) and Peste des petits ruminants virus (PPRV), SARS-CoV-2 (COVID-19) and Climate-aggravated infectious diseases such as those caused by Arboviruses. Under vaccine development, Samuel is employing his immunogenetics expertise to develop anti-east coast fever vaccine by profiling antibody immune (B-cell receptor repertoire) responses to candidate vaccine immunizations. Samuel is also actively involved in developing Genomic Surveillance capacity in Africa.
My Projects
My Publications
Spatiotemporal patterns of Rift Valley fever virus in Africa: a retrospective genomic epidemiology and phylodynamic modelling study

Hyperendemicity of Rift Valley fever in Southwestern Uganda associated with the rapidly evolving lineage C viruses

Unveiling hepatitis E virus diversity in Sudan's internally displaced populations: a molecular epidemiology approach

Methane emissions, productivity parameters, and rumen microbiome across development stages of East African Boran cattle







